Abstract
Proteomic analysis of cells, tissues and body fluids has generated valuable insights into the complex processes influencing human biology. Proteins represent intermediate phenotypes for disease and provide insight into how genetic and non-genetic risk factors are mechanistically linked to clinical outcomes. Associations between protein levels and DNA sequence variants that colocalize with risk alleles for common diseases can expose disease-associated pathways, revealing novel drug targets and translational biomarkers. However, genome-wide, population-scale analyses of proteomic data are only now emerging. Here, we review current findings from studies of the plasma proteome and discuss their potential for advancing biomedical translation through the interpretation of genome-wide association analyses. We highlight the challenges faced by currently available technologies and provide perspectives relevant to their future application in large-scale biobank studies.
Similar content being viewed by others
Introduction
Genome-wide association studies (GWAS) for human diseases have robustly identified large numbers of risk-associated genetic variants implicated in disease susceptibility1. The colocalization of genetic associations for disease traits with those for intermediate molecular phenotypes, such as gene expression2 and metabolomics3, provides powerful evidence to advance hypotheses regarding the genes and pathways through which these disease-associated variants mediate their effects4. Proteins can appear in the blood circulation owing to active secretion or cellular leakage and thereby provide a window into the current state of human health5. Quantitative trait loci (QTLs) for circulating proteins can therefore complement these analyses: collectively, colocalized associations with multiple omic phenotypes offer a route to a comprehensive molecular interpretation of disease GWAS hits6. These analyses have the potential to expose disease-causing pathways, uncover new drug targets, highlight novel therapeutic indications and identify clinically relevant biomarkers7.
However, despite the central importance of proteins in disease pathogenesis, large-scale studies of protein QTLs (pQTLs) have only recently become feasible8,9,10,11,12,13. For most biomedical and clinical applications conducted at a population scale, whole blood and its cell-free derivatives are clinically accessible and minimally invasive specimens that are well suited to the assessment of human health and disease states. Studies of blood-based pQTLs have identified hundreds of associations between single-nucleotide polymorphisms (SNPs) and protein levels6,8,9,10,11,14,15,16,17. Many of these pQTLs colocalize with association signals for common human diseases. Nevertheless, the scale of the largest pQTL studies conducted to date remains limited to the examination of a few thousand proteins across a few thousand individuals. There remains considerable potential in scaling these studies up, in terms of both sample size and proteome coverage.
Beyond the value of large-scale proteomics data to genetic studies, collecting large-scale proteomics data in population studies provides opportunities to study non-genetic associations, making it possible to capture markers of lifestyle and environmental exposure, to stratify individuals according to their state of health (or disease) and to monitor the longitudinal progression of disease. The development of increasingly massive biobanks and population cohorts allows us to envisage deep molecular phenotyping performed across hundreds of thousands of individuals18,19,20,21. These opportunities inevitably raise questions regarding which technologies to prioritize, what outcomes to expect and whether it is worth the investment (given current costs) to characterize the proteomes of entire populations.
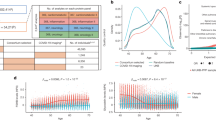
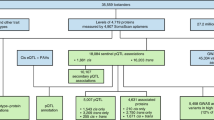
In this Review, we focus on large-scale proteomic technologies currently capable of profiling the blood circulating proteome in large population studies (Fig. 1). We start by reviewing the current state of the field and describe the proteomic composition of serum and plasma. Next, we turn to the experimental challenges related to measuring the circulating proteins, focusing on technologies that are applicable in high throughput to blood-derived samples collected from large population studies. Then we provide an overview of the pQTL studies conducted to date and discuss current challenges and opportunities faced by the field. Finally, we showcase selected applications of pQTL studies in the context of disease GWAS. We close with perspectives for future large-scale studies with proteomics and how the information they provide can be translated into clinical and biomedical applications.
This figure sketches the central steps and challenges encountered when one is conducting a genome-wide association study (GWAS) with proteomics in large population cohorts. a | The blood proteome is a composite of many proteins from different cells and tissues. Some are secreted into the bloodstream to perform some signalling or enzymatic actions, while others may leak from damaged cells and reflect processes occurring elsewhere in the body. b | Mass spectrometry and affinity (antibody-based or aptamer-based) proteomics are the key methods available to determine blood circulating protein levels (Box 1). Each method has its inherent advantages and challenges (Table 1). c | Many approaches used in the statistical analysis of protein quantitative trait locus (pQTL) studies are similar to those applied in other GWAS with high data dimensionality, such as metabolomics. Shown on the left is the minor-allele frequency (MAF), as well as the resultant percentages of homozygous and heterozygous genotypes involving the major (A) and minor (B) alleles. These genotypes are tested for statistical association with protein levels, as shown in the box-and-whisker plot below. As in expression QTL (eQTL) studies, cis and trans associations can be distinguished (middle panel), the latter suggesting possible functional links between the protein encoded at the pQTL locus and the trans-associated proteins (Fig. 2). The resultant pQTLs can be tested for whether they colocalize with disease-associated loci (right panel). d | Newly identified pQTLs should be investigated for potential artefacts, such as epitope effects, cross-reactivity and genotype-dependent binding specificity, cleavage site and peptide mass (see the main text). e | pQTLs have multiple biomedical applications. For instance, pQTLs can be subjected to Mendelian randomization (MR) analysis using proteins as exposures to identify proteins that are causal for disease outcomes (see also Box 2); support drug target selection and validation, as well as drug safety and repurposing; indicate genetic confounders of clinical protein biomarkers (see also Supplementary Table 1); and be integrated in larger systems biology networks to help interpret complex biological processes. The graphic at the top right illustrates the strategy for MR analyses. The graphic at the bottom right shows a hypothetical network where blue nodes are proteins, green nodes are single-nucleotide polymorphisms and red nodes are disease traits. Links between blue and green nodes represent pQTLs, links between red and green nodes represent a disease/trait GWAS hit and links between blue nodes represent correlations between protein levels. FDA, US Food and Drug Administration.
The human plasma proteome
The human proteome is defined by analysis and annotation of all potentially protein-coding genes in the human genome. There are approximately 20,000 genes in humans that provide the blueprint for all proteins to be expressed and processed at any given point in time and in any tissue or organ of the human body. However, translated proteins often undergo further post-translational processing: for example, particular amino acid residues can be modified to modulate their physiological properties through phosphorylation or glycosylation. Most proteins then execute their function in interaction with other partners, either locally in their cells and tissues of origin, or after being transported to act in some distant process. In the following section, we introduce the current efforts of the proteomics community with a focus on approaches and technologies to profile proteins in blood.
Experimental coverage of the human proteome
Critical for any proteomic studies is the definition of the set of proteins that one might expect to detect. As of today, the Human Proteome Project has collected mass spectrometry (MS) data that provide experimental evidence for almost 90% of the approximately 20,000 canonical proteins predicted from genomic open reading frames22. One of the current main efforts of the Human Proteome Project is to find experimental evidence for the predicted — but so far undetected — ‘missing proteins’ and to determine their function. One strategy involves expanding proteomic analyses to rare tissues and cell types under a range of conditions23.
MS remains the most commonly used technology24 in proteomics, alongside complementary approaches using affinity-based assays25,26 or recently emerging protein sequencing methods27. Large-scale MS-based studies have captured nearly 80% of the human proteins observed across several human tissues28,29 and provide access to browse data built from more than 250,000 peptides (see PeptideAtlas, Human Proteome Map and ProteomicsDB in Related links). In parallel, the Human Protein Atlas (HPA) is dedicated to mapping human protein expression across organs and tissues30, including blood31, and to mapping their subcellular localization32. The HPA uses antibodies and transcriptomics to annotate tissue-specific protein expression on a gene-centric level and currently hosts data from 26,000 antibodies targeting proteins from more than 17,000 protein-coding genes (approximately 87% of the human proteome). The HPA has demonstrated that half of all proteins are expressed in all tissues and that instances of tightly restricted protein expression are far less common, except for proteins involved in specialized processes such as spermatogenesis. In terms of their cellular localizations, the HPA found that the products of approximately 13,000 protein-coding genes reside inside cells and the products of approximately 5,500 protein-coding genes reside in membranes, while the products of approximately 2,600 protein-coding genes are secreted into the extracellular space30. Although the proteomic analysis of tissues or cells is less relevant to large-scale population studies, such analyses are central to efforts to understand the mechanisms involved in disease pathogenesis, as well as to put findings from population studies into context.
The plasma proteome
On collection, blood is often centrifuged to generate fractions containing blood cells and a cell-free fluid, which accommodates the circulating proteome. The fluid is called ‘serum’ or ‘plasma’ depending on whether blood clotting is permitted (serum) or inhibited by anticoagulants (plasma) during the blood draw. The key challenges in analysing blood samples are the broad range of concentrations at which circulating proteins appear and the fact that some proteins fluctuate strongly in their abundance in response to disease or other physiological reasons. Collectively, MS or immunoassays have detected about 5,000 circulating proteins, which represent approximately 25% of the human proteome33. Whereas more than 2,300 proteins account for 75% of the overall protein mass in cells, only 20 account for around 90% of the total protein mass in plasma28.
Proteins circulating in plasma can theoretically originate from any organ or cell, or can even pass through the placenta to be exchanged between mother and child34. However, the most abundant proteins, such as albumin, apolipoproteins and components of the complement system, are primarily produced and secreted by the liver. A recent reannotation of the human secretome found that about 730 of the estimated 2,600 secreted proteins have the bloodstream as their primary destination: other secreted proteins can reside locally in the extracellular matrix or in other body fluids35. More information about the human secretome annotation is accessible via the secreted protein section of the HPA portal (see Related links). Proteins secreted or that leak into the circulation are found in a wide range of concentrations and are involved in processes related to blood homeostasis, transportation, defence, signalling, digestion and inhibition of other proteins, inflammation and mechanisms of wound healing35.
Although secretion is the active process directing proteins towards the extracellular space, the presence of cellular proteins in the bloodstream may be a consequence of a variety of different and complex processes, including response to damage to cells and organs (Fig. 1). The detection of leakage proteins, such as cardiac troponin, can provide biomarkers for tissue damage. Knowledge of the tissue and cellular origin of the proteins that leak into the blood can reveal insights into the characteristics of the biosample: proteins leaking from blood cells can be used to assess the quality of sample preparation36.
Current limitations in the knowledge of the circulating proteome
There has been substantial progress in studying the characteristics of proteins across tissues and determining their structure–function relationship in biological systems37. More recently, there have been moves towards single-cell proteomics38. Blood-based proteomics has focused mainly on expanding measures of protein abundance: insights into interactions, structures, isoforms and post-translational modifications of circulating proteins have not been addressed in great detail. Even though these factors contribute to the complexity of identifying and precisely measuring the components of the plasma proteome, current technologies are unable to characterize these at the required throughput, precision and sensitivity. When alternative splicing of transcripts from the approximately 20,000 protein-coding genes is accounted for, the estimated number of possible ‘proteoforms’ reaches 70,000 (ref.39), a number further magnified when all possible combinations of post-translational modifications are considered. Capturing this diversity poses enormous technical challenges considering that different proteoforms of the same protein may coexist in the same sample, and that one of them may be pathogenic, while another may be protective40.
Preanalytical variables can also influence the quantification of circulating proteins, and to minimize possible effects, stringent, consistent and short sample processing procedures are recommended36,41. Variables include the blood collection type (serum or plasma), collection tubes (type of anticoagulants), preservatives (protease inhibitors), separation of blood cells (centrifugation speed), times for preparation (needle to centrifugation) and until storage (needle to freeze), storage format (aliquotation) and biobanking (freezer temperature). Besides inconsistency in plasma purity36, protein degradation occurring after blood draw and over the years of storage can influence the detectable protein levels42. Effects of preanalytical variables have been demonstrated for the processing steps from needle to freezer, such as precentrifugation or postcentrifugation delay43,44,45. In multicentre studies, differences in institutional practices in sample processing, shipping and storage are to be considered. For each proteomics technology or assay used, the influence of preanalytical variables on the protein detectability needs to be considered and assessed46,47,48,49 so that data from different studies can be robustly compared41.
Probing the plasma proteome in high throughput
The ultimate aim of population proteomic analysis is to be able to survey the circulating plasma proteome of individuals at high sensitivity and high specificity combined with a high degree of multiplexing, massive sample throughput and low cost. The challenge lies in applying precise protein-detection methods that are able to quantify thousands of proteins across a wide dynamic range in many thousands of samples, while minimizing the amount of starting material required and the analysis time needed. In the following discussions, we introduce and compare the main technologies that allow the capture of circulating proteins in the context of large-scale population studies; that is, MS and affinity-based methods (for further methodological information, see Box 1).
MS-based proteomics
Since the early years of the first decade of this century, MS-based analysis has been applied to measure circulating proteins5,50. At its core, MS measures the mass-to-charge ratio (m/z) of ionized molecules, such as proteins or peptides, within a gas phase. Although many variations and instruments are used for the ionization, detection, identification and quantification of the ions, plasma proteomics workflows are typically based on an enzymatic digestion of the circulating proteins into peptides. Depletion of highly abundant proteins51 or enrichment of proteins present at low concentrations52 is sometimes applied to address the challenge imposed by the concentration range and dynamics of proteins of the plasma.
There are two complementary approaches to peptide measurement in MS. Targeted MS uses stable-isotope standard peptides as reference points to provide absolute quantification of the peptides in the sample. By contrast, untargeted MS uses the intensity of the peptide ions as a semiquantitative readout of peptide abundance. Particularly for plasma, as peptides representing low-abundance proteins can be masked by peptides originating from more highly abundant proteins, accurate measurement across a wider range of protein abundance concentrations has generally required extensive prefractionation of the sample, limiting MS-based analyses to smaller-scale studies53. Efforts to increase the scale and scope of MS-based assays have revived interest in application to plasma, with recent studies demonstrating reliable detection of approximately 500 abundant proteins in nearly 1,000 samples41,54. In MS studies focusing on detecting as many proteins as possible, up to 4,000 proteins can be identified in a handful of samples55. Although general concepts, advances and the versatility of MS have been reviewed extensively elsewhere37,56, we describe here some recent studies that indicate the growing potential of MS for population-wide studies of the plasma proteome.
For untargeted MS, data-dependent acquisition (DDA) MS methods, which focus on analysing the more prominent peptide ions, have been widely used57. Recent efforts to shorten the analysis time and peptide scanning schemes allow measurement of nearly 300 proteins in 10 plasma samples within 3 hours58, and one recent study included 1,300 sample assays for 450 proteins59. Data-independent acquisition (DIA) approaches consider all peptide ions and are better able to detect less-abundant proteins. DIA requires less complex preanalytical sample handling than DDA but at the cost of increased time for data collection and subsequent bioinformatic analyses to match the extended list of targets with comprehensive protein libraries. One recent DIA-based study described the quantification of 340 proteins in 200 plasma samples from twins60 and revealed that 50% of all detected peptides (so-called peptidoforms) had undergone post-translational modification61. The largest DIA MS proteomics study to date included 1,500 plasma samples, using improved chromatography and increased reproducibility to detect a common set of 320 proteins in more than 90% of the samples62.
For more focused studies, where a smaller number of preselected proteins are of interest, targeted MS workflows can be preferred. In such a setting, targeted MS provides a more sensitive and absolute quantification of protein concentrations. These assays use predetermined analysis conditions and addition of known amounts of synthetic, isotopically labelled peptides (or proteins) to act as references for quantification. This approach facilitates comparison of protein quantities across studies51,63,64. However, the scope of targeted MS is constrained by the predefined sets of labelled canonical peptides offered by companies such as Biognosys (Zurich, Switzerland) or MRM Proteomics (Montreal, Canada).
Affinity-based proteomics
Affinity proteomics has emerged as an attractive alternative to MS-based identification of proteins26 by conducting classical immunoassays at higher throughput and higher sensitivity and in multiplex format25. Among the variety of immunoassays such as ELISA, immunohistochemistry, immunofluorescence, flow cytometry and western blots, all mainly use classic antibodies but may also use alternative binders to detect endogenous proteins in a variety of sample types65. The basic concept for these multianalyte assays66, often making use of advances in DNA technologies (such as DNA microarrays), relies on measuring multiple proteins simultaneously in one sample by miniaturizing the assays. A reduced analytical surface area increases the signal-to-noise ratios by increasing the number of occupied binding sites and avoids the depletion of the analyte in solution.
Over the past two decades, methods for multiplexed protein measurement have emerged to enable analyses across a wide range of concentrations, in thousands of samples. These developments have made these assays attractive for population-wide analyses, and they have been adopted in most recent genome-wide pQTL analyses. These advances in affinity assay methods rely on the improved performance of the binding reagents (antibodies or aptamers) used to detect antigens with high selectivity and high binding affinity67. For analyses of plasma proteins, available platforms range from ultrasensitive single-analyte assays for the detection of circulating proteins at less than 1 pg ml−1 (ref.68) to multiplexed assays that capture more than 4,000 proteins by designed protein-binding DNA aptamers9.
The suspension bead array technology of Luminex (Austin, TX, USA) supports multiplexing via combining different colour-coded microparticles followed by flow cytometry analysis69: each colour code represents one immunoassay for a given protein target. These assays can typically be multiplexed to quantify 30 analytes per sample, processed in batches of up to 384 samples, and have been applied to studies of more than 8,000 samples and up to 50 proteins70. Myriad RBM (Austin, TX, USA) offers services based on a set of more than 300 proteins on the Luminex technology, whereas a variety of predefined and/or customizable panels of sandwich immunoassays are available from providers such as MilliporeSigma, R&D Systems, Bio-Rad and Thermo Fisher. Alternative platforms for multiplexed immunoassays are offered by Quanterix (Billerica, MA, USA), ProteinSimple (San Jose, CA, USA) and Meso Scale Diagnostics (Rockville, MD, USA), and are reviewed elsewhere71.
By contrast, the technology implemented by Olink (Uppsala, Sweden) uses proximity extension assays. Detection of a given protein requires binding of two separate antibodies that carry complementary oligonucleotide tags: when two antibodies bind to the target protein, these oligonucleotides can be hybridized and extended by a DNA polymerase72. This assay therefore uses protein-specific binding properties to generate a readout that then relies on DNA concentrations more easily quantifiable by quantitative PCR. Presently, the technology enables the analysis of up to 1,164 proteins, distributed across 14 themed protein panels, each panel analysing 92 proteins in 90 samples at a time. Recently, one of these panels was deployed across 3,400 individuals to map genetic loci for plasma protein biomarkers in cardiovascular disease73, followed by a meta-analysis in more than 21,000 individuals74.
SomaLogic (Boulder, CO, USA) uses an in-house-developed library of modified aptamers for highly multiplexed protein profiling75. In recent years, its aptamer library has grown from approximately 800 (ref.75) to more than 4,000; in parallel the sample sizes investigated have grown from hundreds to tens of thousands76. The proprietary assay processes 90 samples per batch, and its readout is on DNA arrays. Instead of matching two binding reagents for increased specificity, the selectivity of the platform is built around specific aptamers selected for their capability to bind to their target protein for an extended time (slow off rate).
Comparative discussion of the available methods
Several criteria influence decisions about which technology to deploy for a given large-scale plasma proteome analysis. Although no single method is a universal solution for all analytical aspects, each has particular strengths. There has been no systematic and direct comparison of all available methods, but a starting point for future work could be the ‘popular proteins’77, which is a set of approximately 1,000 circulating proteins that can currently be detected by both MS-based and affinity assay approaches33. Analytical performance criteria for the main technologies are summarized in Table 1, including specificity, reproducibility, sensitivity, degree of multiplexing, sample throughput, quantification and translatability, costs and accessibility of the technology and the derived data.
MS-based approaches benefit from a large community of users and companies. The mode of MS-based protein identification and absolute target quantification as well as future opportunities to capture several post-translational modifications61 are noticeable benefits compared with the affinity assays. However, sample throughput and analytical sensitivity are relatively low (there is no amplification method available), limiting the potential to apply MS-based approaches to large plasma studies. Peptide sequence variants and modifications may further increase the number of missing data points. Newer approaches — such as DIA and variations thereof56 — enable the protein-level data to be digitally stored, so improvements in matching tools to detect peptides together with growing libraries may allow reanalysis of the data in the future.
Affinity-based methods clearly lead on sample throughput (N > 1,000) and analytical sensitivity (below 1 ng ml−1), which can be enhanced by use of signal amplification methods such as are possible with some DNA-based readouts. The number of analytes that can be measured differs between technologies, and several panels may be used to detect large numbers of target proteins. Although the number of reagents to target the human proteome is increasing78, the susceptibility to off-target binding and the lack of reproducibility of assays have raised concerns about the quality of some affinity-based measures79. To achieve higher-quality data, guidelines on the validation of affinity capture-based data have been established80 and systematically applied to a variety of common techniques and samples81,82.
GWAS with proteins circulating in plasma
The goals of proteomic GWAS are to identify genetic sequence variants associated with proteomic features in a given cell or biofluid. Those features are typically protein abundance levels but could extend to other characteristics, such as isoform diversity or post-translational modification. GWAS analysis conducted across the large number of proteomic phenotypes now becoming available for many cohorts presents challenges, not only in terms of computational efforts and data management but also with respect to the quality control of the phenotypic measurements, automation of data analyses, and integration and interpretation of the results.
Identification of pQTL signals
Preprocessing and quality control of proteomic data to avoid spurious association from outliers, or deviations of the protein abundance distributions from assumptions made in standard statistical models, may be cumbersome when one is working with thousands of quantitative traits. Inverse normal scaling of protein levels is often used as a simple and conservative approach to deal with distributional issues and appears to lead to robust associations, as shown by the generally good replication between studies10; however, this is achieved at the cost of some loss in statistical power. Log scaling and/or winsorization — that is, moving extreme values closer to the normal distribution of the proteomic data — offer alternative approaches to inverse normal scaling, and computation of P values based on data preprocessed by multiple methods can be used to obtain consensus associations8. Associations can be computed with the same tools and models as deployed in other GWAS of quantitative traits83,84: batch effects and covariates can be managed by a combination of direct inclusion in analytical models and the use of latent variable approaches, such as PEER85, to ‘find’ the covariates that matter (at some price of removing real effects). Adding specific principal components of the proteomic data as covariates may increase statistical power, for instance where principal components reflect variation induced by processes related to sample handling and storage.
For pQTL discovery studies, the gold standard involves application of a combined genome-wide and proteome-wide significance level together with independent replication. However, this can be a high bar, since a study-wide significance threshold designed to minimize false-positive associations may require researchers to account for the billions of tests performed (a million or more DNA variants for each of several thousand proteins). This approach leaves many studies underpowered to detect genuine but weaker associations. Limiting association testing to variants located in the immediate vicinity of the respective protein-encoding genes can increase statistical power for the detection of cis-pQTLs but restricts the breadth of the study. Novel methods to better incorporate correlative structures between proteins and genetic variants, such as use of sparse multivariate regression models86, may improve this trade-off between power and robustness.
Extending pQTL analyses to include rarer alleles (rather than the common alleles that have been the primary focus of GWAS) may increase opportunities to detect variants with particularly large effects on protein expression and disease risk, since rare alleles are often of more recent origin and may not yet have been subject to negative selection. However, as with disease association studies, the robust detection of rare variant associations can be troublesome, particularly if the rarity of the alleles themselves requires the use of ‘variant aggregation’ tests to provide a signal. Such approaches depend for their power on the ability to combine disparate alleles that have similar broad functional impact, while avoiding dilution of the test by including neutral variants. Such an approach can be difficult enough for coding variants, where the gene provides an obvious unit for aggregation, but is some way from being solved for variants that fall into intergenic and regulatory regions.
pQTL studies to date
Over the past decade, there have been a profusion of GWAS analyses, many also with a limited proteomic scope1,6, including several large-scale studies with multiplexed immunoassays that cover tens of proteins to a few hundred proteins12,13,17,70,73,87. We provide a comprehensive list of all pQTL studies of which we are currently aware in Table 2. To date, only two MS-based studies in blood have combined proteomics with GWAS, one identifying approximately 160 proteins in the plasma of approximately 1,000 individuals53 and the other analysing the small peptide subset of a non-targeted metabolomics platform88. Otherwise, large-scale pQTL studies have made use of affinity proteomics approaches8,9,10,11.
Several of the affinity proteomics studies have used the aptamer approach developed by SomaLogic. The KORA study analysed more than 1,100 proteins in 1,000 participants from a German cohort8 and identified 540 pQTLs that connected 450 independent genetic variants with 280 proteins. This analysis was extended within the INTERVAL cohort of UK blood donors10: an expanded SomaScan panel of almost 3,000 proteins was deployed across 3,300 individuals, raising the pQTL count to 1,930, involving 1,480 proteins. The largest published SomaScan-based pQTL study analysed more than 4,000 proteins across 5,500 Icelanders from the AGES Reykjavik study9 and reported more than 3,130 pQTLs influencing the abundance of 1,800 proteins.
Studies using the Olink platform have tended to include smaller numbers of proteins but larger sample sizes. The most recent analysis, conducted by the SCALLOP consortium, analysed 90 cardiovascular proteins in more than 21,000 individuals74: this study yielded a total of 467 pQTLs influencing 94% of these protein targets.
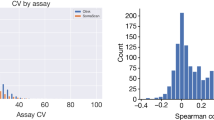
These pQTL studies have provided confidence in the reproducibility of the underlying methods. The three SomaScan studies displayed good agreement: the AGES study confirmed more than 84% of the pQTLs found in the KORA study and more than 72% of those from the INTERVAL study. The INTERVAL study assayed participants with both the SomaScan platform and the Olink platform and found that 65% of the SomaScan-detected pQTLs for overlapping targets were replicated with Olink (with a correlation of 0.95 in effect-size estimates). Some small-scale studies provide data measured in parallel on two platforms89,90. Further comparative studies are needed to fully delineate the consistency between platforms, particularly between MS-based and affinity-based analyses: such analyses have been limited by the sparse overlap between proteins jointly detected on both platforms91.
Cis-pQTLs and trans-pQTLs
Genetic variants that associate with protein levels are generally classified into one of two categories. They are located either in cis, close to the gene that encodes the associated protein, or in trans, at greater distance, typically on a different chromosome (Fig. 2). Between 18% and 25% of the proteins assayed by aptamer assays have been found, at current sample sizes, to have a significant cis-pQTL. Twenty-seven per cent of the pQTLs reported by the KORA study8 were trans-pQTLs, rising to 70% in the larger INTERVAL study10. The greater multiple testing burden inherent in trans-pQTL analysis, together with the increased potential for non-genetic phenotypic variability (Fig. 2), makes sample size a critical factor in trans-pQTL detection.
A protein quantitative trait locus (pQTL) is a locus containing one or more genetic variants that associate with protein levels. Genetic variants can affect many biological processes in the life cycle of a protein, which eventually leads to their association with protein levels measured in a blood sample. Genetic variants can act in cis, that is, affect processes involved in the expression of the associated protein itself (blue), or act in trans, and modify the associated protein through various indirect processes (red). Variants can directly change the properties of a protein or mediate their effects through regulatory feedback loops that affect protein levels. Variants can also exert regulatory effects at the RNA level, such as by altering transcriptional output of mRNAs (for instance by changing DNA methylation sites or altering transcription factor binding), modifying RNA structure, altering mRNA splicing, impacting RNA turnover or affecting ribosomal binding. pQTLs may also act through more complex processes, such as affecting co-translational protein folding or microRNA (miRNA) expression or target specificity. Some of the variation in protein levels may also reflect differential experimental response to changes in protein epitopes. Identification of the functional variants of a pQTL can therefore be challenging.
Cis-pQTLs indicate the presence of a variant that is likely to have a direct and causal effect on the observed protein levels at that locus. If the causal variant of a cis-pQTL acts primarily through mRNA expression or turnover, an (RNA) expression QTL (eQTL) may also be found in a relevant tissue or cell type. If instead, the cis variant alters protein abundance through an impact on protein translation or turnover, a cis-eQTL is less likely, but it may still be observed if expression is upregulated through compensatory feedback to maintain protein levels. Although the presence of a nearby eQTL generally supports the identification of a causal pQTL variant, it is also possible that the two signals map close together by chance or that the eQTL and pQTL arise from different variants that are in high linkage disequilibrium. A variant in a shared promoter of two genes may further complicate the picture, giving rise to more complex hypotheses regarding the link between the variant and the associated protein level.
Trans-pQTLs are of particular inferential value because a trans-pQTL implies an interaction between an — often still to be identified — causal gene at the pQTL locus and the associated protein encoded at the trans position, pointing to novel pathways of protein regulation or interaction. Sun et al.10 present several examples including the identification of PRDM1 as the probable causal gene at an inflammatory bowel disease GWAS locus, and the dissection of complex association signals for antineutrophil cytoplasmic antibody-associated vasculitis at the SERPINA1 locus. Orthogonal evidence based on pharmacological intervention and transgenic mice can support hypothesized causal gene-to-protein relationships from trans-pQTLs, as exemplified by the recent study by the SCALLOP consortium on eight gene products targeted by compounds or antibodies in clinical development74. Genetic loci involving multiple cis-pQTLs and trans-pQTLs are particularly informative, as they link multiple proteins in putative gene–protein networks that can suggest new hypotheses as to their possible interactions and functions. A genetic variant associated with a trans-pQTL can further change protein levels in trans through regulation or modification of cis-encoded microRNAs or epigenetic marks. The study by the SCALLOP consortium identified 30 trans-pQTLs that involved two or more proteins, with ABO, ST3GAL4, JMJD1C, SH2B3 and ZFPM2 showing association with the levels of 5 or more of the 90 proteins analysed by one panel of the Olink platform74.
Colocalization with eQTLs
Most disease GWAS signals map to regulatory rather than coding sequences, and the downstream effectors through which they operate are typically unclear92. Evidence of enrichment between patterns of disease association and the location of tissue-specific regulatory sequences (such as enhancers), as well as for the sites of tissue-specific eQTLs, indicates that the identification of colocalizing cis-eQTLs has value in highlighting those effectors and reconstructing disease mechanisms93. However, it is also clear that this approach is not as robust as has often been assumed: at many loci, there are multiple colocalizing cis-eQTLs, implicating several candidate effectors, and it seems intrinsically unlikely that all are directly mediating disease risk94. pQTL analysis provides a complementary approach to reconstructing links between genetic variation, molecular processes and disease predisposition. In recent studies8,9,10, only 40% of detected cis-pQTLs could be shown to colocalize with cis-eQTLs detected in projects such as Genotype–Tissue Expression (GTEx)2, which generated cis-eQTL data for approximately 40 tissues in up to 900 individuals. It remains to be seen how the recently reported contamination of GTEx data with highly expressed, tissue-enriched genes impacts on this observation95.
The discrepancy between pQTL and eQTL discovery has led to questions about their relative merits, questions which, by and large, ignore the fact that these data types are quite distinct. Most obviously, large-scale pQTL data are almost entirely restricted to the circulating proteome, whereas projects such as GTEx have provided eQTL maps for multiple tissues: there will be many loci where the downstream effectors (for example, transcription factors) are not represented in the circulation and where any contribution from pQTL studies will need to await technologies that provide tissue-specific, and even cell-specific, proteomic readouts. On the other hand, there will be other loci where the mediating mechanisms can be detected only through pQTL analyses: where the genetic variants exert their effects through an impact on protein stability or modification, for example, or where the protein half-life in the circulation far exceeds that of the RNA.
Data sharing
Sharing of full summary statistics — that is, reporting all pairwise variant–protein associations with effect size and standard error regardless of their significance levels — can be data storage intensive but is an essential component of scientific discourse. The GWAS catalogue1 now accepts and maintains full summary association statistics data sets, and summary statistics for some of the larger pQTL studies are already freely available8,10. Access to summary data enables immediate incorporation into online tools for causal inference that use Mendelian randomization approaches, such as EpiGraphDB96 and MRbase97. Whereas sharing of summary-level data should be a prerequisite for publication, proactive sharing of individual-level data (both raw and processed genotypes and protein measurements) renders data sets far more valuable facilitators of scientific discovery, fostering development of novel statistical approaches and maximizing the opportunities for integration with other complementary sources of data. Controlled-access repositories, such as EGA and dbGAP, provide a mechanism to support the sharing of sensitive individual-level data with bona fide investigators.
Working with ratios
It is possible to conduct association analyses focused not only on the expression levels of individual proteins but also on the ratios between them. Inspired by the use of hypothesis-free testing of metabolite ratios in metabolomic GWAS98, the challenge with ratio-based analyses is the massive explosion in the number of statistical tests that could be performed and the consequent need to allow for the inflation in type 1 error that could result. However, as with metabolomic data, where known relationships between products and substrates can be used to constrain the scope of ratio testing (avoiding testing of all pairwise combinations), there may be opportunities to perform protein-ratio QTL analyses in subsets of functionally connected proteins. For example, the strength of the pQTL association of rs41341749 with CCL14 and CCL23 increased by 35 orders of magnitude when the ratios of their abundance were used, an effect replicated in an independent cohort8. A gain in the strength of association when ratios are used may also result when one of the proteins concerned is a proxy for variability induced by sample handling and storage, or any other common normalizing factor.
Network integration
A broader view of disease biology can be enabled by analyses that present the relationships between proteins as networks, especially when those networks can be integrated with additional information such as that provided by genetic associations and colocalization of those associations with clinical end points. Network relationships between proteins can be derived from existing biological knowledge, using databases such as WikiPathways99 and STRING100, or they can emerge from data-driven integration in multidimensional omics data sets. For example, Gaussian graphical models (GGMs) have been shown to reconstruct pathway reactions from high-throughput metabolomics data101, and, when integrated with metabolic and disease GWAS associations, are powerful tools to mine high-dimensional GWAS data sets for biomedically relevant associations102. GGMs that include proteomic data recently revealed a genome–proteome–disease subnetwork implicated in Crohn’s disease8.
One illustrative example involves the pleiotropicABO locus. In GWAS, this locus has been associated with numerous disease outcomes, including coronary artery disease103 and venous thromboembolism104. This locus includes at least three independent, widely replicated genetic signals that influence the plasma abundance of multiple proteins in trans8,9,10,14,91,105. Two of these signals tag the blood group B and O alleles, and the third lies upstream of the ABO gene. A network based around these trans proteins, assembled from literature findings, experimental protein–protein interactions and the partial correlations between their abundances, indicates that these proteins play a joint role in angiogenesis and that this angiogenic function is modulated by variation at the three ABO signals8. Further analyses of such network models are required to establish the functional relevance of observed links between GGM network nodes, possibly using statistical approaches to objectively define functional modules in such networks106.
Analytical considerations
As with all high-throughput omics technologies, proteomic data are subject to multiple potential sources of error and bias, which the user must consider to avoid misinterpretation or overinterpretation of the results (Table 1).
Epitope effects
One challenge in interpreting cis-pQTL associations from affinity assay-based GWAS is the possibility that a genetic variant in the protein-coding sequence modifies the binding epitope of the protein that is recognized by an assay’s antibody or aptamer, in the absence of other biological consequences14. Such an epitope effect can be introduced by a non-synonymous mutation within the target epitope. Alternatively, it may reflect larger-scale changes in the protein structure arising from variants (including frameshift insertions or deletions) that modify the protein’s overall structure or folding, or its potential for post-translational modification. Linkage disequilibrium (that is, correlation) between such a variant influencing epitope binding and otherwise entirely unrelated variants within adjacent non-coding sequence can result in a pattern of association that is indistinguishable from a genuine cis-pQTL.
There are several ways to establish whether epitope effects could be driving an observed cis-pQTL signal8,9,10,14. Evidence that the gene implicated in a cis-pQTL contains a coding variant (which may require interrogation of imputed or sequenced whole-genome sequence data) raises this as a concern: the coding variant needs to be evaluated for its impact on protein sequence and structure. Experimentally, one could produce both protein variants and quantify differential recognition by directly comparing assay performance. The presence of a colocalizing eQTL can be reassuring, serving as a pointer that the observed differences in protein abundance are likely to result from altered protein expression rather than differential recognition. However, this is not fail-safe: the eQTL could itself be an artefact if the coding variant leads to allelic bias in the mapping of RNA sequencing reads. Where pQTLs are the consequence of SNP-associated allele-specific gene expression, RNA sequencing can be used to provide evidence that only one protein isoform is expressed. Replication of a pQTL on multiple platforms, or when a range of binders and assays are used, provides confidence that the association reflects changes in protein levels rather than binding affinity. pQTLs that display exceptionally large effect sizes and that show no colocalization with other GWAS phenotypes are more likely to represent epitope effects, as large differences in protein levels might be expected to have measurable biological impact.
Trans-acting epitope effects are also possible. This can happen when a coding variant at a pQTL signal influences the properties of a protein-modifying enzyme or an interaction partner in cis: this can affect the epitopes of proteins modified by this enzyme or those which interact in a genotype-dependent manner with the binding partner8. In some relevant loci, binders have been selected explicitly to bind different epitopes of the same target: the use of multiple SomaScan aptamers to target the products of the various APOE alleles is one such example8.
Protein-altering variants can also influence MS-derived data107. If a SNP introduces a new protein cleavage site (or eliminates an expected one), the peptides generated by enzymatic digestion may differ from those listed in the canonical reference peptide library. A possible workaround for capturing these instances is to apply a ‘proteogenomics’ approach108, in which peptide variants are imputed from the genetic data and then used to build a variant-aware peptide reference109.
From the findings taken together, it is clear that epitope effects represent a challenge to the interpretation of pQTL studies and require careful annotation of the assays used. However, these effects also constitute opportunities for further analyses of biological variation in the protein sequences that have no direct impact on the biological function (until shown otherwise). It is important to bear in mind that pQTLs identified with affinity assays after all represent genetic differences in the amount of affinity-captured proteins, which is then merely interpreted as representing protein abundances.
Binding specificity and cross-reactivity
Affinity-based assays rely on the correct identification of their intended targets, but cross-reactivity and lack of specificity have been a concern81. The selectivity and suitability of affinity reagents are context and technology dependent and hence must be evaluated for each sample type and application71. For example, a SomaScan-based proteome analysis of blood from young and old parabiotic mice reported an age-related reduction in circulating levels of growth/differentiation factor 11 (GDF11), raising the possibility that restoring normal levels of GDF11 could reverse ageing. However, it was subsequently established that the assay used to identify and isolate GDF11 had low specificity110. Cross-reactivity to similar epitopes, such as those encoded by paralogous genes, or domains shared across different proteins may confound an association signal10. Cross-target binding may also occur when binders designed to target non-human proteins are included in the panel, such as viral or bacterial proteins on the SomaScan platform3. In the absence of the intended target, pQTLs identified with such binders may correspond to genuine genetic signals of another, yet unidentified target protein. Finally, genuine errors in binder selection, for instance due to misidentification of the intended target protein, constitute another potential source of error.
Binding specificity has been addressed by several studies8,9,10,14. For instance, Sun et al.10 assessed potential off-target cross-reactivity for 920 aptamers and found that 14% showed comparable binding with a homologous protein, nearly half of which were alternative forms of the same protein. Emilsson et al.9 provided evidence for target specificity for 773 affinity binders for a panel of 4,137 aptamers by using affinity pull-down followed by MS-based target identification, although only some of the experiments were conducted with blood serum or plasma. Recently, SomaLogic conducted a systematic analysis of the reliability and specificity of SOMAmer protein-affinity reagents in the SomaScan assay. They found that out of 1,612 tested SOMAmer reagents, 73% did not bind to any related proteins, 14% bound to related proteins with at least tenfold weaker affinity and 13% bound to other unrelated proteins with similar affinity. They further confirmed specific target enrichment by pull-downs from human plasma for 123 of the SOMAmer reagents111. A recent article from SomaLogic also provides updated information on MS-target confirmation and characterization of possible cross-binding and off-target effects for many of the proteins on its in-house platform76.
The use of community standards for validation80 and open access to validation data will help to drive improvements: it should be possible to understand, for any given binder, not only that it captures the intended target but also which other circulating proteins can be enriched. Sharing raw data from multiplatform studies may allow independent evaluation of platform performance89. The integration of such information into public protein databases with cross-referencing identifiers to affinity proteomics platforms should increase the accuracy of pQTL inference.
Clinical and biomedical applications
The characterization of pQTLs drives various biomedical and pharmaceutical applications. pQTLs provide intermediate phenotypes to interpret the findings of disease GWAS, clues to genes that are causal for disease biology, opportunities to discover clinical biomarkers, matches between existing drugs and new disease indications, pointers to potential safety concerns for drugs in development, insights into protein–protein interaction networks and much more (Box 2). Here, we discuss some of the key applications in further detail.
Genetic variance in clinical biomarker proteins
The measurement of protein levels in accessible biofluids (for example, plasma, urine or cerebrospinal fluid) is one of the mainstays of clinical medicine, providing many biomarkers with diagnostic or prognostic value. The ability to generate robust pQTL associations at scale could not only provide mechanistic insights into disease biology that prioritize targets for therapeutic development but could also promote the discovery of novel clinical biomarkers. Some of these biomarkers will support more accurate diagnosis of disease, whereas others will increase the detection of drug side effects, stratify disease risk or act as surrogates of developing disease that are valuable in clinical trials.
Many frequently used clinical tests are blood based; nearly half involve the measurement of proteins54. Historically, these tests have involved measurement of the abundance of single proteins, but multiprotein biomarkers are emerging, one example being the profiling of cardiovascular risk in patients with coronary heart disease112. The current set of clinically used proteins represents only a fraction of the circulating proteome, suggesting considerable opportunities to develop improved protein signatures for monitoring human health and disease states76. Ten years ago, Anderson113 identified 109 US Food and Drug Administration (FDA)-approved protein analytes that can be assayed in serum or plasma. An update of this list (Supplementary Table 1) now includes 199 unique analytes. Among the subset of these FDA-approved protein analytes for which assays were available in at least one of the three recent large-scale SomaScan studies8,9,10, almost one-third have pQTLs with an effect size large enough to be detected (Table 3). This observation, which is in agreement with recent reports of the impact of ancestry on protein biomarker levels114, indicates that some of the observed differences in disease prevalence between ethnicities may have a genetic basis reflected in blood protein abundances and that reference intervals for those biomarkers should be tailored to ancestry.
Interpretation of the findings of disease GWAS
One of the key motivations behind large-scale pQTL analyses is to support efforts to relate protein abundance levels to the growing inventory of disease-associated genetic variants and thereby to accelerate the identification of potentially translatable biomarkers. As the scale and scope of pQTL studies have expanded, so too has the list of pQTLs coincident with disease risk variants8,9,10. In recent SomaScan articles8,9, between 11.5% and 20.7% of cis-pQTLs were in high linkage disequilibrium (r2 ≥ 0.8) with sentinel disease-associated variants. Webservers such as PhenoScanner115 and SNiPA116 integrate GWAS hits from multiple sources and can be used to identify and interpret such overlapping signals. By addition of data from large-scale studies of RNA expression, such as those provided by GTEx, it becomes possible to construct causal chains leading from DNA to RNA and on to protein and disease2. Tissue-specific protein expression data gathered from the HPA30 can further illuminate these pathways. One important caveat is that the rapidly growing numbers of GWAS associations and pQTLs mean that some of the apparent overlap in genomic locations is merely the consequence of two distinct signals — driven by entirely different sequence variants — which happen to map to the same stretch of DNA but which have no mechanistic relationship. It is therefore essential to confirm that coincident signals are driven by the same genetic variants (that is, that they colocalize) before assuming any biological connection117.
Polygenic scores
One powerful strategy for biomarker discovery is emerging from recent advances in the use of polygenic risk scores to stratify disease risk across populations118. These scores aggregate the information on individual genetic predisposition for a given disease across many thousands of risk variants. For many common conditions, these scores can identify individuals who, on the basis of their patterns of shared common sequence variation, are at substantially increased (or decreased) future risk of disease118. Proteomic analyses conducted in individuals from these extremes of disease risk, ideally using plasma samples collected from healthy individuals many years ahead of clinical disease onset, offer a route to accelerate the discovery of prognostic biomarkers. It is possible to directly evaluate relationships between disease polygenic risk scores and protein expression levels in cross-sectional cohorts using a Mendelian randomization framework119. As an example of the power of such an approach, Mosley et al.120 intersected coronary artery disease risk score data with an aptamer-based pQTL study of 759 individuals, computing a ‘virtual proteome’ for each individual, which was then evaluated for its association with clinical end points in a much larger cohort.
To the extent that these biomarkers capture processes fundamental to disease risk, they have the potential to profile individual risk irrespective of its basis — genetic or otherwise — in the same way that cholesterol levels synthesize both genetic and lifestyle-related contributions to the risk of coronary disease. There are opportunities to extend these approaches beyond measures of overall disease risk to characterize biomarkers that capture the relative contribution of multiple disease-associated processes to disease progression in a given individual. This information can be parlayed into improved tools for prognostication and therapeutic optimization. In the case of type 2 diabetes, for example, it is possible to construct a set of ‘partitioned’ risk scores, each made up of variants that influence one of several pathways contributing to disease risk (such as obesity, fat distribution, insulin resistance and insulin secretion)121 and then to use them in the search for process-specific biomarkers.
Inferring causality
Although pQTL and eQTL analyses can generate important hypotheses linking a disease risk variant to a plausible downstream effector, these inferences are correlative rather than causal. Even in the ideal setting, whereby a disease risk variant is shown to colocalize with a pQTL for a protein with a biologically plausible link to disease, there is no formal proof that the protein lies on the causal pathway to disease. Instead, variation in the circulating levels of that protein might be just one of several molecular events attributable to the risk variant, only one of which is causally linked to disease development. Alternatively, variation in the biomarker may be secondary to, rather than causal for, the disease process (‘reverse causation’). Such pQTLs may still point to useful biomarkers of disease predisposition or severity, but interventions that restore levels of the biomarker to normal levels will not necessarily be disease modifying. In the case of coronary artery disease, for example, the evidence indicates that triglycerides and LDL cholesterol play a causal role, but the levels of HDL cholesterol and C-reactive protein do not.
Proof of a causal connection ultimately relies on demonstrating — in a suitable system and using a disease-relevant readout — that perturbation of the expression or function of the protein of interest has a material effect on the development of disease. Fortunately, there are a host of approaches for examining the consequences of protein perturbation, ranging from cellular screens, through manipulation in animal models, to the detection, in isolated populations or those with high levels of consanguinity, of individuals who have inherited genotypes that result in extreme (high or low) levels of the protein. Here too, the integration of genetic and proteomic data can be extremely valuable. Finding that coding variants in the gene encoding a protein of interest are causally implicated with disease provides reassurance that a regulatory pQTL involving the same gene is also likely to be causal. Similarly, evidence from Mendelian randomization studies that a given protein is the target of several distinct cis-pQTLs and trans-pQTLs, each of which also colocalizes with a disease GWAS signal, represents consistent evidence that the protein is mediating disease risk rather than simply reflecting the activity of another causal process with which it happens to share some regulatory overlap122.
Mendelian randomization analyses are central to many of these approaches, providing what can be considered the genetic equivalent of a randomized clinical trial. For example, large-scale SomaScan analyses have implicated several protein pathways as causal for disease risk, including a protective role for prostate secretory protein of 94 amino acids (PSP94) in prostate cancer, and associations between raised IL1RL2 and IL18R1 levels and atopic dermatitis risk, and between higher MMP12 levels and decreased risk of coronary disease risk and stroke9. Yao and colleagues13 used pQTL data to identify six proteins likely to be causal for coronary heart disease, two of which, cystatin C and PON1, were also associated prospectively with death from new-onset coronary heart disease or cardiovascular disease in the Framingham Heart Study. Chong and colleagues123 conducted a systematic Mendelian randomization meta-analysis of the circulating proteome based on pQTL data from more than 20,000 individuals8,9,10,70,73 and disease GWAS data from up to 400,000 participants. This study revealed seven causal mediators for ischaemic stroke, including a protective role for SCARA5 and TNFSF12, highlighting important roles for stroke protein biomarkers in cardiovascular and non-cardiovascular diseases and singling out LPA, F11 and SCARA5 as particularly attractive drug targets. Many recent association studies with intermediate proteomics traits include Mendelian randomization analyses11,74,87,124, and new methods are emerging to cope with the inherent challenges of a multi-omics Mendelian randomization approach, such as pleiotropy between the traits.
Conclusions and perspectives
Genetic association studies using intermediate phenotypes represent analyses of experiments conducted by nature, able to provide a valuable resource of biological information. Proteomic data at scale have only recently become accessible to large-scale GWAS approaches, largely due to analytical limitations that are specific to the composition of the plasma proteome. Advances in affinity proteomics technologies are rapidly remedying this shortfall, with recent studies reporting the parallel measurement of 5,000 proteins in almost 17,000 blood samples76, and many substantially larger studies are under way74. Currently, between 10% and 20% of all pQTLs discovered were found to colocalize with clinical GWAS loci. This overlap can be expected to increase as the scale and scope of both pQTL studies and clinical GWAS continues to increase. This will add to the value of plasma proteomic data especially for the biomedical and pharmaceutical field by providing more and better instruments for drug target validation7 and causal inference96. Crucially, large-scale pQTL studies in large healthy cohorts complement the growing use of plasma proteomic data in the context of case–control datasets and in clinical trials, improving inference in study designs that are subject to reverse causation and confounding.
Although affinity assays lead the way in pQTL mapping, it is wise to remain aware of their limitations. Associations of interest should be supported by independent validation of target specificity and should include consideration of possible cross-reactivity and epitope effects. As none of the current methods is the optimal or sole solution, validation across multiple technologies will be key, with the most important findings further supported by cellular and other functional studies. It remains to be shown how feasible MS-based approaches will be for large-scale GWAS projects. Although low sample throughout and low analytical sensitivity currently limit the use of MS-based techniques in population studies, MS has the potential to identify a broader range of peptides and proteins, as well as a variety of post-translational modifications and splice forms. GWAS of IgG glycosylation125 and total N-glycans126 have already reported biologically relevant associations, most of them with proteins that are involved in protein glycosylation. Other approaches, known as ‘glycoproteogenomics’, seek to understand the differential glycosylation of plasma protein biomarkers127. Future efforts should consequently also attempt to capture the different modification characteristics of proteins by high-throughput and quantitative technologies and possibly determine tissue-specific isoforms that are of relevance for these traits. In parallel, the prospects of combining data from large-scale proteomics with molecular readouts, such as DNA methylation87,124,128,129, metabolomics130 and glycomics131, opens new avenues for studying human health at a molecular level in population-scale studies.
Recent advances in the scale and scope with which it is possible to survey the proteomic content of plasma and other biofluids are allowing proteomics to take its place alongside the comprehensive characterization possible for other omics approaches, such as those focused on genetic variation and RNA expression. These advances open new opportunities to use proteomics to deliver improved understanding of the mechanistic basis of disease132,133 and to promote novel translational strategies through target and biomarker identification74,76.
References
MacArthur, J. et al. The new NHGRI-EBI catalog of published genome-wide association studies (GWAS catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Lonsdale, J. et al. The genotype-tissue expression (GTEx) project. Nat. Genet. 45, 580–585 (2013).
Suhre, K. et al. Human metabolic individuality in biomedical and pharmaceutical research. Nature 477, 54–60 (2011).
Kastenmuller, G., Raffler, J., Gieger, C. & Suhre, K. Genetics of human metabolism: an update. Hum. Mol. Genet. 24, R93–R101 (2015).
Anderson, N. L. & Anderson, N. G. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell. Proteomics 1, 845–867 (2002).
Melzer, D. et al. A genome-wide association study identifies protein quantitative trait loci (pQTLs). PLoS Genet. 4, e1000072 (2008).
Plenge, R. M., Scolnick, E. M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Suhre, K. et al. Connecting genetic risk to disease end points through the human blood plasma proteome. Nat. Commun. 8, 14357 (2017). This is one of the first GWAS using the SomaScan platform for 1,100 proteins.
Emilsson, V. et al. Co-regulatory networks of human serum proteins link genetics to disease. Science 361, 769–773 (2018). This is currently the largest GWAS using the updated SomaScan platform for 4,000 proteins and 4,000 samples.
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018). This is a recent GWAS using the SomaScan platform with 3,000 proteins on 3,000 samples.
Benson, M. D. et al. Genetic architecture of the cardiovascular risk proteome. Circulation 137, 1158–1172 (2018).
Zhernakova, D. V. et al. Individual variations in cardiovascular-disease-related protein levels are driven by genetics and gut microbiome. Nat. Genet. 50, 1524–1532 (2018).
Yao, C. et al. Genome-wide mapping of plasma protein QTLs identifies putatively causal genes and pathways for cardiovascular disease. Nat. Commun. 9, 3268 (2018).
Enroth, S., Johansson, A., Enroth, S. B. & Gyllensten, U. Strong effects of genetic and lifestyle factors on biomarker variation and use of personalized cutoffs. Nat. Commun. 5, 4684 (2014). This is an early GWAS using the Olink platform; the study highlights the potential impact of epitope effects on protein readouts.
Lourdusamy, A. et al. Identification of cis-regulatory variation influencing protein abundance levels in human plasma. Hum. Mol. Genet. 21, 3719–26 (2012).
Sasayama, D. et al. Genome-wide quantitative trait loci mapping of the human cerebrospinal fluid proteome. Hum. Mol. Genet. 26, 44–51 (2017).
Sun, W. et al. Common genetic polymorphisms influence blood biomarker measurements in COPD. PLoS Genet. 12, e1006011 (2016).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018). This study highlights the potential of large biobanks.
German National Cohort (GNC) Consortium. The German National Cohort: aims, study design and organization. Eur. J. Epidemiol. 29, 371–82 (2014).
Precision Medicine Initiative (PMI) Working Group Report to the Advisory Committee to the Director, NIH. The Precision Medicine Initiative Cohort Program – Building a Research Foundation for 21st Century Medicine (National Institutes of Health, 2015).
Chen, Z. et al. China Kadoorie Biobank of 0.5 million people: survey methods, baseline characteristics and long-term follow-up. Int. J. Epidemiol. 40, 1652–1666 (2011).
Omenn, G. S. et al. Progress on identifying and characterizing the human proteome: 2018 metrics from the HUPO Human Proteome Project. J. Proteome Res. 17, 4031–4041 (2018).
Baker, M. S. et al. Accelerating the search for the missing proteins in the human proteome. Nat. Commun. 8, 14271 (2017).
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Stoevesandt, O. & Taussig, M. J. Affinity proteomics: the role of specific binding reagents in human proteome analysis. Expert. Rev. Proteom. 9, 401–14 (2012).
Smith, J. G. & Gerszten, R. E. Emerging affinity-based proteomic technologies for large-scale plasma profiling in cardiovascular disease. Circulation 135, 1651–1664 (2017).
Timp, W. & Timp, G. Beyond mass spectrometry, the next step in proteomics. Sci. Adv. 6, eaax8978 (2020).
Kim, M. S. et al. A draft map of the human proteome. Nature 509, 575–81 (2014).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Uhlen, M. et al. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Uhlen, M. et al. A genome-wide transcriptomic analysis of protein-coding genes in human blood cells. Science 366, eaax9198 (2019).
Thul, P. J. et al. A subcellular map of the human proteome. Science 356, eaal3321 (2017).
Schwenk, J. M. et al. The human plasma proteome draft of 2017: building on the Human Plasma PeptideAtlas from mass spectrometry and complementary assays. J. Proteome Res. 16, 4299–4310 (2017). This article reviews recent advances in plasma proteomics and uses data from the community to summarize the circulating proteins detected by MS.
Pernemalm, M. et al. In-depth human plasma proteome analysis captures tissue proteins and transfer of protein variants across the placenta. Elife 8, e41608 (2019).
Uhlen, M. et al. The human secretome. Sci Signal 12, eaaz0274 (2019). This article reviews the actively secreted proteins of the human proteome for their destination and reveals that only approximately 730 proteins are secreted into the circulation.
Geyer, P. E. et al. Plasma proteome profiling to detect and avoid sample-related biases in biomarker studies. EMBO Mol. Med. 11, e10427 (2019).
Aebersold, R. & Mann, M. Mass-spectrometric exploration of proteome structure and function. Nature 537, 347–55 (2016).
Marx, V. A dream of single-cell proteomics. Nat. Methods 16, 809–812 (2019).
Aebersold, R. et al. How many human proteoforms are there? Nat. Chem. Biol. 14, 206–214 (2018).
Theodoratou, E. et al. The role of glycosylation in IBD. Nat. Rev. Gastroenterol. Hepatol. 11, 588–600 (2014).
Ignjatovic, V. et al. Mass spectrometry-based plasma proteomics: considerations from sample collection to achieving translational data. J. Proteome. Res. 18, 4085–497 (2019).
Enroth, S., Hallmans, G., Grankvist, K. & Gyllensten, U. Effects of long-term storage time and original sampling month on biobank plasma protein concentrations. EBioMedicine 12, 309–314 (2016).
Kofanova, O. et al. IL8 and IL16 levels indicate serum and plasma quality. Clin. Chem. Lab. Med. 56, 1054–1062 (2018).
Qundos, U. et al. Profiling post-centrifugation delay of serum and plasma with antibody bead arrays. J. Proteom. 95, 46–54 (2013).
Daniels, J. R. et al. Stability of the human plasma proteome to pre-analytical variability as assessed by an aptamer-based approach. J. Proteome. Res. 18, 3661–3670 (2019).
Kim, C. H. et al. Stability and reproducibility of proteomic profiles measured with an aptamer-based platform. Sci. Rep. 8, 8382 (2018).
Shen, Q. et al. Strong impact on plasma protein profiles by precentrifugation delay but not by repeated freeze-thaw cycles, as analyzed using multiplex proximity extension assays. Clin. Chem. Lab. Med. 56, 582–594 (2018).
Di Girolamo, F., Alessandroni, J., Somma, P. & Guadagni, F. Pre-analytical operating procedures for serum low molecular Weight protein profiling. J. Proteom. 73, 667–77 (2010).
Zimmerman, L. J., Li, M., Yarbrough, W. G., Slebos, R. J. & Liebler, D. C. Global stability of plasma proteomes for mass spectrometry-based analyses. Mol. Cell. Proteomics 11, M111.014340 (2012).
Shen, Y. et al. Characterization of the human blood plasma proteome. Proteomics 5, 4034–45 (2005).
Abbatiello, S. E. et al. Large-scale interlaboratory study to develop, analytically validate and apply highly multiplexed, quantitative peptide assays to measure cancer-relevant proteins in plasma. Mol. Cell. Proteomics 14, 2357–74 (2015).
Harney, D. J. et al. Small-protein enrichment assay enables the rapid, unbiased analysis of over 100 low abundance factors from human plasma. Mol. Cell. Proteomics 18, 1899–1915 (2019).
Johansson, A. et al. Identification of genetic variants influencing the human plasma proteome. Proc. Natl Acad. Sci. USA 110, 4673–8 (2013).
Geyer, P. E., Holdt, L. M., Teupser, D. & Mann, M. Revisiting biomarker discovery by plasma proteomics. Mol. Syst. Biol. 13, 942 (2017).
Keshishian, H. et al. Multiplexed, quantitative workflow for sensitive biomarker discovery in plasma yields novel candidates for early myocardial injury. Mol. Cell. Proteomics 14, 2375–93 (2015).
Ludwig, C. et al. Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Mol. Syst. Biol. 14, e8126 (2018).
Doerr, A. Mass spectrometry-based targeted proteomics. Nat. Methods 10, 23 (2013).
Geyer, P. E. et al. Plasma proteome profiling to assess human health and disease. Cell Syst. 2, 185–95 (2016).
Geyer, P. E. et al. Proteomics reveals the effects of sustained weight loss on the human plasma proteome. Mol. Syst. Biol. 12, 901 (2016).
Liu, Y. et al. Quantitative variability of 342 plasma proteins in a human twin population. Mol. Syst. Biol. 11, 786 (2015).
Rosenberger, G. et al. Inference and quantification of peptidoforms in large sample cohorts by SWATH-MS. Nat. Biotechnol. 35, 781–788 (2017).
Bruderer, R. et al. Analysis of 1508 plasma samples by capillary-flow data-independent acquisition profiles proteomics of weight loss and maintenance. Mol. Cell. Proteomics 18, 1242–1254 (2019).
Addona, T. A. et al. Multi-site assessment of the precision and reproducibility of multiple reaction monitoring-based measurements of proteins in plasma. Nat. Biotechnol. 27, 633–41 (2009).
Percy, A. J. et al. Method and platform standardization in MRM-based quantitative plasma proteomics. J. Proteom. 95, 66–76 (2013).
Stoevesandt, O. & Taussig, M. J. Affinity reagent resources for human proteome detection: initiatives and perspectives. Proteomics 7, 2738–50 (2007).
Ekins, R. P. Multi-analyte immunoassay. J. Pharm. Biomed. Anal. 7, 155–68 (1989).
Ayoglu, B. et al. Systematic antibody and antigen-based proteomic profiling with microarrays. Expert Rev. Mol. Diagn. 11, 219–34 (2011).
Rissin, D. M. et al. Single-molecule enzyme-linked immunosorbent assay detects serum proteins at subfemtomolar concentrations. Nat. Biotechnol. 28, 595–9 (2010).
Fulton, R. J., McDade, R. L., Smith, P. L., Kienker, L. J. & Kettman, J. R. Jr. Advanced multiplexed analysis with the FlowMetrix system. Clin. Chem. 43, 1749–56 (1997).
Ahola-Olli, A. V. et al. Genome-wide association study identifies 27 loci influencing concentrations of circulating cytokines and growth factors. Am. J. Hum. Genet. 100, 40–50 (2017).
Fredolini, C. et al. Immunocapture strategies in translational proteomics. Expert Rev. Proteom. 13, 83–98 (2016).
Assarsson, E. et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 9, e95192 (2014).
Folkersen, L. et al. Mapping of 79 loci for 83 plasma protein biomarkers in cardiovascular disease. PLoS Genet. 13, e1006706 (2017).
Folkersen, L. et al. Genomic evaluation of circulating proteins for drug target characterisation and precision medicine. Preprint at bioRxiv https://doi.org/10.1101/2020.04.03.023804 (2020). This is currently one of the largest pQTL studies, with more than 21,000 samples on a 92-protein panel from the Olink platform.
Gold, L. et al. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5, e15004 (2010).
Williams, S. A. et al. Plasma protein patterns as comprehensive indicators of health. Nat. Med. 25, 1851–1857 (2019).
Lam, M. P. et al. Data-driven approach to determine popular proteins for targeted proteomics translation of six organ systems. J. Proteome Res. 15, 4126–4134 (2016).
Colwill, K. & Graslund, S. A roadmap to generate renewable protein binders to the human proteome. Nat. Methods 8, 551–8 (2011).
Baker, M. Reproducibility crisis: blame it on the antibodies. Nature 521, 274–6 (2015).
Uhlen, M. et al. A proposal for validation of antibodies. Nat. Methods 13, 823–7 (2016).
Fredolini, C. et al. Systematic assessment of antibody selectivity in plasma based on a resource of enrichment profiles. Sci. Rep. 9, 8324 (2019).
Edfors, F. et al. Enhanced validation of antibodies for research applications. Nat. Commun. 9, 4130 (2018).
Aulchenko, Y. S., Ripke, S., Isaacs, A. & van Duijn, C. M. GenABEL: an R library for genome-wide association analysis. Bioinformatics 23, 1294–6 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–75 (2007).
Stegle, O., Parts, L., Piipari, M., Winn, J. & Durbin, R. Using probabilistic estimation of expression residuals (PEER) to obtain increased power and interpretability of gene expression analyses. Nat. Protoc. 7, 500–7 (2012).
Ruffieux, H., Davison, A. C., Hager, J. & Irincheeva, I. Efficient inference for genetic association studies with multiple outcomes. Biostatistics 18, 618–636 (2017).
Ahsan, M. et al. The relative contribution of DNA methylation and genetic variants on protein biomarkers for human diseases. PLOS Genet. 13, e1007005 (2017).
de Vries, P. S. et al. Whole-genome sequencing study of serum peptide levels: the Atherosclerosis Risk in Communities study. Hum. Mol. Genet. 26, 3442–3450 (2017).
Graumann, J. et al. Multi-platform affinity proteomics identify proteins linked to metastasis and immune suppression in ovarian cancer plasma. Front. Oncol. 9, 1150 (2019).
Billing, A. M. et al. Complementarity of SOMAscan to LC-MS/MS and RNA-seq for quantitative profiling of human embryonic and mesenchymal stem cells. J. Proteom. 150, 86–97 (2017).
Ruffieux, H. et al. A Bayesian joint pQTL study sheds light on the genetic architecture of obesity. Preprint at bioRxiv https://doi.org/10.1101/524405 (2019).
Freedman, M. L. et al. Principles for the post-GWAS functional characterization of cancer risk loci. Nat. Genet. 43, 513–8 (2011).
Gamazon, E. R. et al. Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation. Nat. Genet. 50, 956–967 (2018).
Wainberg, M. et al. Opportunities and challenges for transcriptome-wide association studies. Nat. Genet. 51, 592–599 (2019).
Nieuwenhuis, T. O. et al. Consistent RNA sequencing contamination in GTEx and other data sets. Nat. Commun. 11, 1933 (2020).
Zheng, J. et al. Phenome-wide Mendelian randomization mapping the influence of the plasma proteome on complex diseases. Preprint at bioRxiv https://doi.org/10.1101/627398 (2019).
Hemani, G. et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife 7, e34408 (2018).
Petersen, A. K. et al. On the hypothesis-free testing of metabolite ratios in genome-wide and metabolome-wide association studies. BMC Bioinformatics 13, 120 (2012).
Slenter, D. N. et al. WikiPathways: a multifaceted pathway database bridging metabolomics to other omics research. Nucleic Acids Res. 46, D661–D667 (2018).
Szklarczyk, D. et al. The STRING database in 2017: quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368 (2017).
Krumsiek, J., Suhre, K., Illig, T., Adamski, J. & Theis, F. J. Gaussian graphical modeling reconstructs pathway reactions from high-throughput metabolomics data. BMC Syst. Biol. 5, 21 (2011).
Shin, S. Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46, 543–550 (2014).
van der Harst, P. & Verweij, N. Identification of 64 novel genetic loci provides an expanded view on the genetic architecture of coronary artery disease. Circ. Res. 122, 433–443 (2018).
Klarin, D., Emdin, C. A., Natarajan, P., Conrad, M. F. & Kathiresan, S. Genetic analysis of venous thromboembolism in UK Biobank identifies the ZFPM2 locus and implicates obesity as a causal risk factor. Circ. Cardiovasc. Genet. 10, e001643 (2017).
Nath, A. P. et al. Multivariate genome-wide association analysis of a cytokine network reveals variants with widespread immune, haematological, and cardiometabolic pleiotropy. Am. J. Hum. Genet. 105, 1076–1090 (2019).
Do, K. T., Rasp, D. J. N., Kastenmuller, G., Suhre, K. & Krumsiek, J. MoDentify: phenotype-driven module identification in metabolomics networks at different resolutions. Bioinformatics 35, 532–534 (2019).
Zhang, B. et al. Proteogenomic characterization of human colon and rectal cancer. Nature 513, 382–7 (2014).
Nesvizhskii, A. I. Proteogenomics: concepts, applications and computational strategies. Nat. Methods 11, 1114–25 (2014).
Ting, Y. S. et al. PECAN: library-free peptide detection for data-independent acquisition tandem mass spectrometry data. Nat. Methods 14, 903–908 (2017).
Harper, S. C. et al. Is growth differentiation factor 11 a realistic therapeutic for aging-dependent muscle defects? Circ. Res. 118, 1143–50 (2016).
SomaLogic. Short Technical Note: Characterization of the Binding Specificity of SOMAmer Reagents in the SomaScan Assay (2019).
Ganz, P. et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA 315, 2532–41 (2016).
Anderson, N. L. The clinical plasma proteome: a survey of clinical assays for proteins in plasma and serum. Clin. Chem. 56, 177–85 (2010). This is an early survey that lists the FDA-approved plasma biomarkers (an update of this list is provided in Supplementary Table 1).
Sjaarda, J. et al. Influence of genetic ancestry on human serum proteome. Am. J. Hum. Genet. 106, 303–314 (2020).
Staley, J. R. et al. PhenoScanner: a database of human genotype-phenotype associations. Bioinformatics 32, 3207–3209 (2016).
Arnold, M., Raffler, J., Pfeufer, A., Suhre, K. & Kastenmuller, G. SNiPA: an interactive, genetic variant-centered annotation browser. Bioinformatics 31, 1334–6 (2015).
He, X. et al. Sherlock: detecting gene-disease associations by matching patterns of expression QTL and GWAS. Am. J. Hum. Genet. 92, 667–80 (2013).
Khera, A. V. et al. Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat. Genet. 50, 1219–1224 (2018).
Richardson, T. G., Harrison, S., Hemani, G. & Davey Smith, G. An atlas of polygenic risk score associations to highlight putative causal relationships across the human phenome. Elife 8, e43657 (2019).
Mosley, J. D. et al. Probing the virtual proteome to identify novel disease biomarkers. Circulation 138, 2469–2481 (2018).
Udler, M. S. et al. Type 2 diabetes genetic loci informed by multi-trait associations point to disease mechanisms and subtypes: A soft clustering analysis. PLoS Med. 15, e1002654 (2018).
Plump, A. & Davey Smith, G. Identifying and validating new drug targets for stroke and beyond. Circulation 140, 831–835 (2019).
Chong, M. et al. Novel drug targets for ischemic stroke identified through mendelian randomization analysis of the blood proteome. Circulation 140, 819–830 (2019).
Hillary, R. F. et al. Genome and epigenome wide studies of neurological protein biomarkers in the Lothian Birth Cohort 1936. Nat. Commun. 10, 3160 (2019).
Shen, X. et al. Multivariate discovery and replication of five novel loci associated with immunoglobulin G N-glycosylation. Nat. Commun. 8, 447 (2017).
Sharapov, S. Z. et al. Defining the genetic control of human blood plasma N-glycome using genome-wide association study. Hum. Mol. Genet. 28, 2062–2077 (2019).
Lin, Y. H., Zhu, J., Meijer, S., Franc, V. & Heck, A. J. R. Glycoproteogenomics: a frequent gene polymorphism affects the glycosylation pattern of the human serum fetuin/alpha-2-HS-glycoprotein. Mol. Cell. Proteomics 18, 1479–1490 (2019).
Zaghlool, S. B. et al. Epigenetics meets proteomics in an epigenome-wide association study with circulating blood plasma protein traits. Nat. Commun. 11, 15 (2020).
Huan, T. et al. Genome-wide identification of DNA methylation QTLs in whole blood highlights pathways for cardiovascular disease. Nat. Commun. 10, 4267 (2019).
Zaghlool, S. B. et al. Deep molecular phenotypes link complex disorders and physiological insult to CpG methylation. Hum. Mol. Genet. 27, 1106–1121 (2018).
Suhre, K. et al. Fine-mapping of the human blood plasma n-glycome onto its proteome. Metabolites 9 (2019).
Gudmundsdottir, V. et al. Circulating protein signatures and causal candidates for type 2 diabetes. Diabetes https://doi.org/10.2337/db19-1070 (2020).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Kim, S. et al. Influence of genetic variation on plasma protein levels in older adults using a multi-analyte panel. PLoS ONE 8, e70269 (2013).
Kauwe, J. S. et al. Genome-wide association study of CSF levels of 59 Alzheimer’s disease candidate proteins: significant associations with proteins involved in amyloid processing and inflammation. PLoS Genet. 10, e1004758 (2014).
Deming, Y. et al. Genetic studies of plasma analytes identify novel potential biomarkers for several complex traits. Sci. Rep. 6, 18092 (2016).
Solomon, T. et al. Associations between common and rare exonic genetic variants and serum levels of 20 cardiovascular-related proteins: the Tromso study. Circ. Cardiovasc. Genet. 9, 375–83 (2016).
Di Narzo, A. F. et al. High-throughput characterization of blood serum proteomics of ibd patients with respect to aging and genetic factors. PLoS Genet. 13, e1006565 (2017).
Carayol, J. et al. Protein quantitative trait locus study in obesity during weight-loss identifies a leptin regulator. Nat. Commun. 8, 2084 (2017).
Solomon, T. et al. Identification of common and rare genetic variation associated with plasma protein levels using whole-exome sequencing and mass spectrometry. Circ. Genom. Precis. Med. 11, e002170 (2018).
Sliz, E. et al. Genome-wide association study identifies seven novel loci associating with circulating cytokines and cell adhesion molecules in Finns. J. Med. Genet. 56, 607–616 (2019).
Gilly, A. et al. Whole genome sequencing analysis of the cardiometabolic proteome. Preprint at bioRxiv https://doi.org/10.1101/854752 (2020).
Orru, V. et al. Genetic variants regulating immune cell levels in health and disease. Cell 155, 242–56 (2013).
Patin, E. et al. Natural variation in the parameters of innate immune cells is preferentially driven by genetic factors. Nat. Immunol. 19, 302–314 (2018).
Acknowledgements
K.S. is supported by the Biomedical Research Program at Weill Cornell Medicine in Qatar, a programme funded by the Qatar Foundation. J.M.S. is supported by the KTH Center for Applied Precision Medicine funded by the Erling Persson Family Foundation and acknowledges the Knut and Alice Wallenberg Foundation for funding the Human Protein Atlas. J.M.S. and M.I.M. acknowledge the Innovative Medicines Initiative Joint Undertaking under grant agreement no. 115317 (DIRECT), the resources of which are composed of a financial contribution from the European Union’s Seventh Framework Programme and an EFPIA companies’ in kind contribution. The views expressed in this article are those of the authors and not necessarily those of the UK NHS, the UK NIHR, the UK Department of Health or the Qatar Foundation.
Author information
Authors and Affiliations
Contributions
K.S. and J.M.S. researched data for article. All authors contributed to the discussion of content, writing the article and reviewing/editing the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
M.I.M. has served on advisory panels for Pfizer, NovoNordisk and Zoe Global, has received honoraria from Merck, Pfizer, NovoNordisk and Eli Lilly, has stock options in Zoe Global and has received research funding from AbbVie, AstraZeneca, Boehringer Ingelheim, Eli Lilly, Janssen, Merck, NovoNordisk, Pfizer, Roche, Sanofi Aventis, Servier and Takeda. As of June 2019, M.I.M. is an employee of Genentech and holds stock in Roche. K.S. and J.M.S. declare no competing interests.
Additional information
Peer review information
Nature Reviews Genetics thanks M. Altelaar and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
A table of all published GWAS with proteomics: http://www.metabolomix.com/a-table-of-all-published-gwas-with-proteomics/
Human Plasma Proteome Project: https://www.hupo.org/plasma-proteome-project
Human Proteome Map: http://www.humanproteomemap.org
In vitro test systems that have been categorized by the FDA: https://www.fda.gov/medical-devices/medical-device-databases/clinical-laboratory-improvement-amendments-download-data
MRbase: a database and analytical platform for Mendelian randomization: http://www.mrbase.org
neXtProt, a knowledgebase on human proteins: https://www.nextprot.org
PeptideAtlas: http://www.peptideatlas.org
pGWAS server: connecting genetic risk to disease end points through the human blood plasma proteome: http://proteomics.gwas.eu
PhenoScanner: a database of human genotype-phenotype associations: http://www.phenoscanner.medschl.cam.ac.uk
ProteomeXchange, a consortium to provide coordinated data submission and dissemination of proteomics repositories: http://www.proteomexchange.org
ProteomicsDB: https://www.proteomicsdb.org
SNiPA: a tool for annotating and browsing genetic variants: http://snipa.org
SomaLogic white paper on SOMAmer specificity: https://somalogic.com/technology/our-platform/somamer-specificity/
The Genome Aggregation Database (gnomAD) browser: https://gnomad.broadinstitute.org/
The Genotype-Tissue Expression (GTEx) project portal: https://gtexportal.org
The Human Protein Atlas: https://www.proteinatlas.org
The Human Protein Atlas: the proteins actively secreted to human blood: https://www.proteinatlas.org/humanproteome/blood/secreted+to+blood
The NHGRI-EBI catalogue of published genome-wide association studies: https://www.ebi.ac.uk/gwas
Uniprot Proteomes — Homo sapiens: https://www.uniprot.org/proteomes/UP000005640
Supplementary information
Glossary
- Colocalization
-
Two genetic associations are said to be colocalized if the strengths of their statistical associations covary at a genetic locus, suggesting a shared genetic causal variant for the observed associations.
- Protein QTLs
-
(pQTLs). A protein quantitative trait locus (pQTL) is an association of protein levels at a genetic locus; it is often represented by the strongest associating single-nucleotide polymorphism.
- pQTL studies
-
Genome-wide association studies where the dependent variables are the levels of proteins measured using a proteomics approach. The identified loci that associate with protein levels are termed ‘protein quantitative trait loci’ (pQTLs).
- Open reading frames
-
Portions of DNA that can be translated into protein and that are terminated by a stop codon.
- Post-translational modifications
-
Biochemical modification of the primary peptide sequence, typically by covalent addition of a chemical group, such as for phosphorylation and glycosylation. Post-translational modifications can change the accessibility to a protein epitope and potentially influence the binding of affinity reagents.
- Data-dependent acquisition
-
(DDA). A data acquisition mode used in mass spectrometry analysis where only a selected set of peptides with the most intense peptide ions are being fragmented and analysed.
- Data-independent acquisition
-
(DIA). A data acquisition mode used in mass spectrometry analysis where all peptides detected within a particular window of the mass-to-charge ratio are being fragmented and analysed.
- Aptamers
-
Short single-stranded (and possibly modified) nucleotides that are selected from a synthetic library of sequences to recognize a specific target protein (for example, via structural elements) with high affinity.
- cis-pQTLs
-
When a protein quantitative trait locus (pQTL) is at or near the genetic locus that encodes the associated protein; often an ad hoc distance cut-off is used to differentiate cis-pQTLs from trans-pQTLs. A cis-pQTL suggests a direct influence of a genetic variant at that locus on protein expression or turnover.
- trans-pQTLs
-
When a protein quantitative trait locus (pQTL) is distant from the protein-coding gene or on another chromosome. A trans-pQTL indicates an indirect link between the genetic locus and protein expression or turnover.
- Linkage disequilibrium
-
Two genetic loci are in linkage disequilibrium if their genotypes correlate within a population. Lack of recombination between loci results in them commonly being co-inherited as a haplotype.
- Mendelian randomization
-
A method to estimate the unconfounded effect of an exposure (for example, protein level) on an outcome (for example, disease risk) using genetic variation.
- Gaussian graphical models
-
(GGMs). Network representations of the partial correlations between a set of quantitative variables, here the protein levels. Partial correlations used in a protein GGM can be viewed as the amount of pairwise correlation between the levels of two proteins that remains when the contributions of all other proteins are accounted for.
- Pleiotropic
-
A genetic locus is pleiotropic when one or more of its variants is associated with two or more seemingly unrelated phenotypic traits.
- Epitope effect
-
An effect of an epitope-changing variant on the binding properties of affinity reagents with regard to their antigens. A difference in reported antigen recognition may be mistaken for a difference in protein abundance.
- Polygenic risk scores
-
Combined risk scores derived from a weighed combination of genetic associations, possibly including millions of associations.
Rights and permissions
About this article
Cite this article
Suhre, K., McCarthy, M.I. & Schwenk, J.M. Genetics meets proteomics: perspectives for large population-based studies. Nat Rev Genet 22, 19–37 (2021). https://doi.org/10.1038/s41576-020-0268-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41576-020-0268-2